Methods of tracking bass.
The idea of following, or tracking, fish as they make their way around their unseen watery world has always held a fascination for anglers. This is not without significant challenges due to the dynamic nature of the ocean, the uncertain and invisible movement of individuals across huge ranges, and the changes we are currently witnessing in our ocean habitats. The science in this area is moving ahead, with several exciting and fascinating techniques being used in addition to tagging, and others constantly being worked on. BASS members have been involved in several projects in this field, and we are continually looking for new ones to support.
Tagging
Mark-Recapture
Traditional methods of tagging fish involve Mark-Recapture (MR) tags. These identify where the tagged fish ends up, allowing its journey between release and recapture to be inferred. BASS members who participated in the major CEFAS tagging exercise in the early 2000’s will be familiar with these. MR tags may also be used to estimate fish population size.
Of course, fish don’t travel in straight lines; their journey may be more complex than can be discerned by the use of MR tags. These are still valuable, but in more recent times tags are being used which can actually follow fish as they move around. These include:
Data Storage Tags These record many parameters, including the depth (pressure), temperature, tidal movements, salinity, light and magnetic field experienced by the fish on a regular basis, to produce individual fish histories. The chosen parameters are fed into algorithms to construct tracks showing the wide-scale movements of each fish. Such tags as these were used in the C-BASS project. Ewan Hunter from CEFAS gave a presentation at the 2016 BASS AGM C-BASS presentation 1 C-BASS presentation 2
Acoustic Telemetry tracks fish within a fixed area, like an estuary, using small acoustic transmitter tags, which emit a unique “ping” approximately every 90 seconds. These pings can be detected by series of strategically placed acoustic receivers. Such tags negate the need to recapture individuals, or recover the tags, in order to produce data. BASS was involved in the I-BASS project run by Dr Tom Stamp, which used this type of tag.
To see a short video about acoustic telemetry, click here.
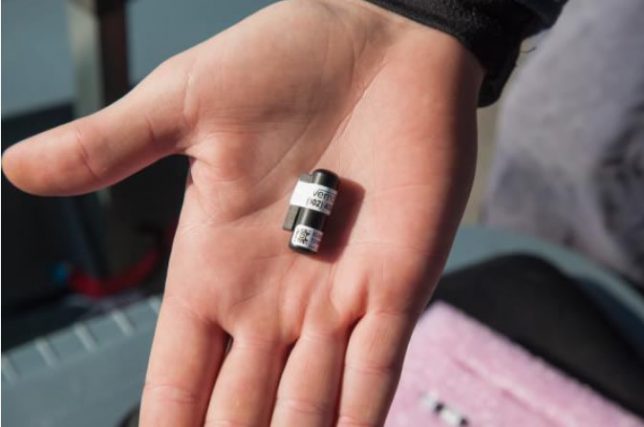
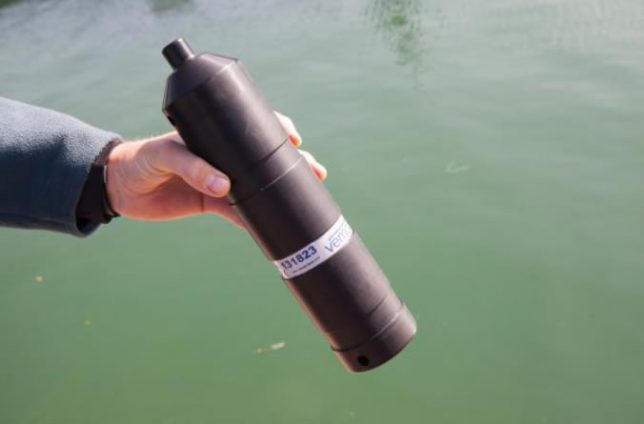


With any tag there are potential downsides, such as higher rates of infection and inflammation where tags are attached, possible expulsion of the tags, and altered behaviour due to swimming/feeding performance rendering inaccurate results from individuals. The extent of any problems is proportional to the size of the tag.
Chemical analysis
Otolith microchemical analysis
Who would have thought that the ear bones, or otoliths, of a fish could reveal important information about its life history. These natural tags are sometimes referred to as the ‘black box recorder’ of the fish. Studying isotopes of carbon, nitrogen, oxygen and strontium found in this metabolically inert environmental recorder provides an accurate way to analyse stock structures, migration patterns, spawning areas/natal origin (area of birth), nursery areas and dietary patterns – even years later.
The chemical composition found within discrete layers of the otolith reflects that of the ambient water in which the fish is swimming, and its diet, at the time each layer was laid down. These discrete layers create a permanent temporal record of the fish’s movements and diet.
One drawback to the use of otoliths is the uncertainties in stable isotope values across the ocean basin. Terrestrially, isotope composition varies in a predictable fashion, creating isoscapes researchers use as accurate geospatial tags for land dwelling species. The difficulty in marine environments is the lack of predictability due to the highly dynamic habitat, however technology is developing to produce accurate measurements of oceanic isotopic compositions.
Another negative is the need to kill the fish to obtain otoliths. Scales, which avoid this, can be used for microchemical analysis, but may not give such good results since they only provide data from the most recent season, due to the constant turnover of material.
Refs: Wikipedia
Chemical archives in fishes beyond otoliths
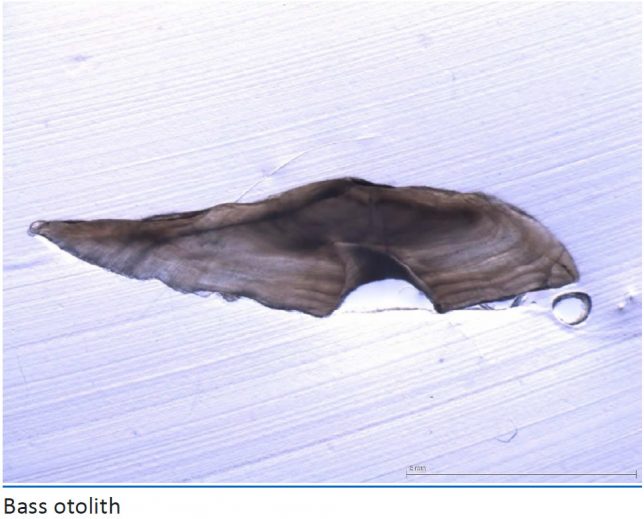 Image from CEFAS Report of the Seabass (Dicentrarchus labrax) Otolith and Scale Exchange Scheme 2011
Image from CEFAS Report of the Seabass (Dicentrarchus labrax) Otolith and Scale Exchange Scheme 2011
See also C-BASS presentation 1 & 2 above.
Environmental DNA analysis
Fish (and other organisms) leave traces of themselves via the loss of DNA into their surroundings from scales, faeces, mucus, eggs etc. Incredibly, it is possible to detect and analyse minute quantities of this in the aquatic environment.
This so-called environmental DNA, or eDNA, can be analysed by DNA sequencing; DNA metabarcoding compares the results with previously studied DNA libraries to identify individual, predetermined species. This technique allows biomonitoring using water samples without having to collect and identify the living organism, which can be difficult with traditional methods, for example when sampling larvae in spawning areas.
Problems with using eDNA include difficulty in determining the origin of DNA, e.g. where was individual when it left the trace, particularly in coastal areas. More research is needed to determine the spatial and temporal preservation of DNA. eDNA analysis has also been shown to calculate biomass in lab studies, but not as of yet in the wild, however in time it could be used as passive means of estimation stock size.
Ref Wikipedia and Environmental DNA
BASS Science Group, Nov 2020.
With thanks to Dr Tom Stamp, University of Plymouth and Joe Dawson, University